This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison
|
Conclusions
|
|
Introduction
Nijmegen Breakage Syndrome, a rare disease of chromosomal instability, is a life altering disorder [1]. With no know cure and few treatment options aimed at only a subset of its side effects, research surrounding NBS is in need of expansion. Because the overwhelming majority of patients with this disease die due to predisposition and development of cancer, the primary goal of the specific aims presented here is to determine the molecular mechanism behind the regulation of NBN, the mutated protein in NBS patients [2]. Better understanding these mechanisms could prove beneficial in attempts to prevent cancer development in NBS patients and ultimately anyone who may suffer from NBN-related cancer predisposition. Gathered information from past research has shown that NBN interacts with two other proteins, MRE11 and RAD50, to form the MRN complex. Involved in sensing and repairing DNA double-strand breaks (DSB), mediating DSB-related cell cycle checkpoint signaling and telomere maintenance, the complex, including NBN, is considered a tumor suppressor, preventing the accumulations of errors leading to death or uncontrolled growth of the cell [3]. Because it is known that the NBN expression levels are negatively correlated with cancer predisposition in NBS patients, understanding the molecular mechanisms that regulate its expression may offer answers for cancer prevention and treatment [4]. Humans and mice have shown to possess similar phenotypes, including cancer predisposition, making them a great model for NBS research. Additionally, the use of naked mole-rats, a rodent with innately low cancer susceptibility, may offer additional information regarding NBN in relation to cancer [5]. Using genomic and proteomic techniques, I will test my hypothesis that the expression levels of NBN are critical indicators of cancer susceptibility, not only in NBS patients, but in all organisms. Aim 1
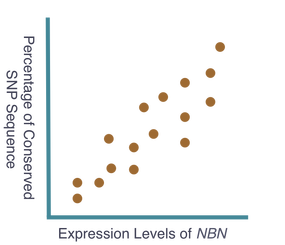
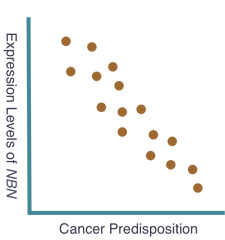
The goal of my first aim is to identify any single nucleotide polymorphisms (SNPs) directly affecting the expression levels of NBN. To accomplish this, I will sample of population of individuals, gathering NBN expression levels and sequenced genomes from each. The individual sequenced genome, obtained through next generation sequencing, will be compared to the corresponding expression level of NBN, obtained using quantitative mass spectrometry. Subsequently, these data will be compared among all individuals in the tested population and studied for correlation. I hypothesize that a correlation will exist and individuals with high expression of NBN will have shared SNPs that indicate potential regulatory sequences for NBN expression. Moreover, individuals with these shared SNPs will possess similar cancer predisposition. These expected results can be seen below. The ability to assess these SNPs and determine their role in the expression of NBN may offer a potential area of research resulting in the manipulation of NBN expression levels and therefore manipulation of cancer predisposition. Additionally, the sampled population will be continually observed to gather information on individual cancer development. These data will be added to the previous data to further support the original hypothesis that NBN expression levels are critical indicators of cancer predisposition.
Aim 2
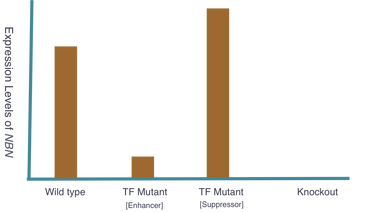
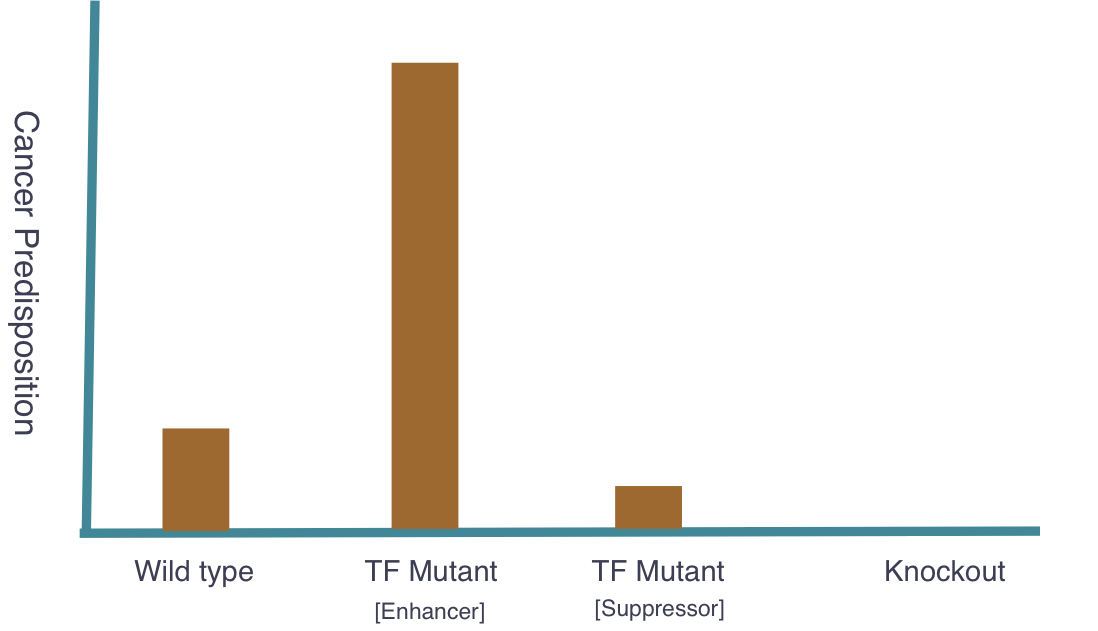
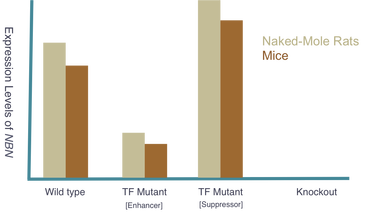
The goal of my second aim is to identify transcription factors (TFs) that regulate NBN expression. Using mice, I hypothesize that there exists specific TFs that affect NBN expression and therefore cancer susceptibility. Three groups of mice will be used and the following outcomes are expected:
Long-term cancer screenings of these mice groups will further support the original hypothesis that NBN expression levels are critical indicators of cancer predisposition.
Aim 3
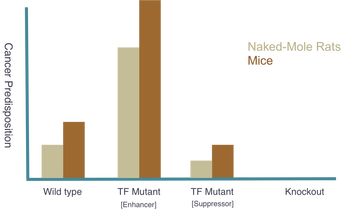
The goal of my third aim is to identify NBN expression levels in naked mole-rats and compare and apply the findings to Mice. I hypothesize that naked-mole rats will express an innately higher amount of NBN, which will be confirmed through quantitative mass spectrometry. Additionally, I will replicate the three groups used in Aim 2 (Wild type, Knockout, and TF mutant) with the naked mole-rats to observe any similarities or differences in comparison to other organisms. Expected results can be seen below. Because of their low susceptibility to cancer, naked mole-rats pose as a great model for cancer research. Proving they possess naturally higher levels of NBN would further support the hypothesis that NBN expression levels are critical indicators of cancer predisposition. As in the other two aims, these organisms will be observed for cancer development and results will be compared to their personal expression levels. Findings are expected to further support the original hypothesis that NBN expression levels are critical indicators of cancer predisposition.
Future Directions
Presumably, I will have proven the negative correlation between expression of NBN and cancer predisposition. With this knowledge and the identification of SNPs and TFs affecting the regulation of NBN, we may refocus research to target these mechanisms and regulators. Additional research surrounding small molecules that could alter NBN expression would be advantageous to those affected by NBS. With a long-term goal to help those affected by NBS, we aim to provide information in helping lead to new treatment options, such as gene therapy and drug developments, to ultimately influence the levels of NBN and ultimately cancer predisposition. Additionally, understanding how to manipulate levels of NBN could prove beneficial to the entire population as screening for individuals with innately low expression of this gene/protein could aid in cancer prevention. |
| |||||||||||||
|
References:
[1] Nijmegen Breakage Syndrome. (2011, April 1). Retrieved January 30, 2015, from http://ghr.nlm.nih.gov/condition/nijmegen-breakage-syndrome [2] Chrzanowska, K., Janniger, C. (2014, Septemper 29). Nijmegen Breakage Syndrome. Retrieved January 30, 2105, from http://emedicine.medscape.com/article/1116869-overview [3] Lamarche, B., Orazio, N., & Weitzman, M. (2010). The MRN complex in double-strand break repair and telomere maintenance. FEBS Letters, 3682-3695. Retrieved March 13, 2015, from http://www.ncbi.nlm.nih.gov/pmc/articles/PMC2946096/ [4] Maser, R., Zinkel, R., & Petrini, J. (2001). An alternative mode of translation permits production of a variant NBS1 protein from the common Nijmegen breakage syndrome allele. Nature Genetics, 27. Retrieved March 13, 2015, from http://www.nature.com/ng/journal/v27/n4/abs/ng0401_417.html [5] Buffenstien, R. (2005). The Naked Mole-Rat: A New Long-Living Model for Human Aging Research. The Journals of Gerontology Series A: Biological Sciences and Mecdical Sciences, 1369-1377. Retrieved March 19, 2015 from http://biomedgerontology.oxfordjournals.org/content/60/11/1369.full.pdf+html |