This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison
|
Domains
What is a protein domain?
Domains are conserved structural units of a protein that typically have a structural, functional or interacting role in the protein's overall function [1]. A protein may have a single domain, multiple domains, many of the same domain or a variety of different domains. A domain is not specific to any single protein. Domains play a roles in [2]:
|
Click proteins to enlarge
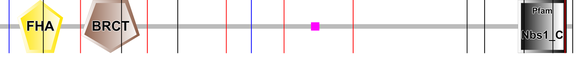
Figure 1. NBN_HUMAN protein domains as determined by SMART. The small pink square represents an area of low complexity.
Figure 2. NBN_HUMAN protein domains as determined by Pfam. The red domain represents the Nbs1_C domain while the small blue rectangle represents an area of low complexity and the grey regions represent disorder areas.
|
NBN Protein Domains
|
NBN Protein Homologs
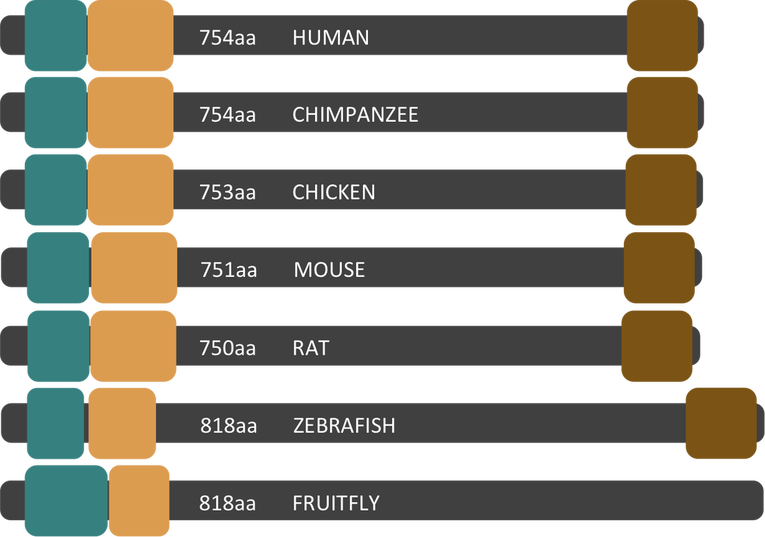
Figure 3. Domains of NBN homologs across seven select model organisms.
|
Motifs
What is a motif?
A motif is a short, conserved region that is presumed to have a biological function [1]. These regions can be molecular sequences, such as nucleotides or amino acids, or they can be structural units, such as a alpha-helix or beta-sheet. Motifs can be seen throughout domains. |
NBN Protein Motifs
|
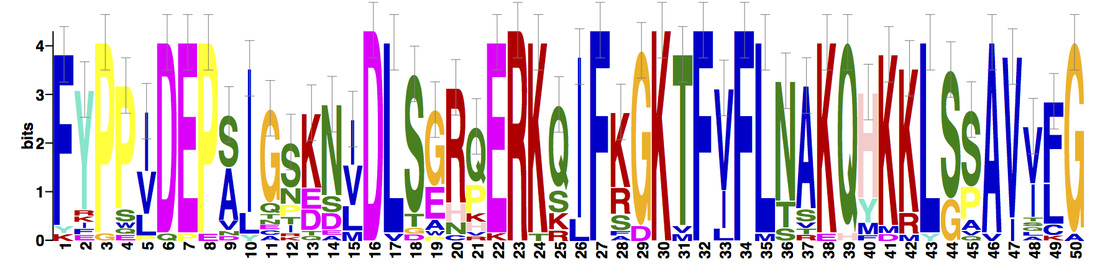
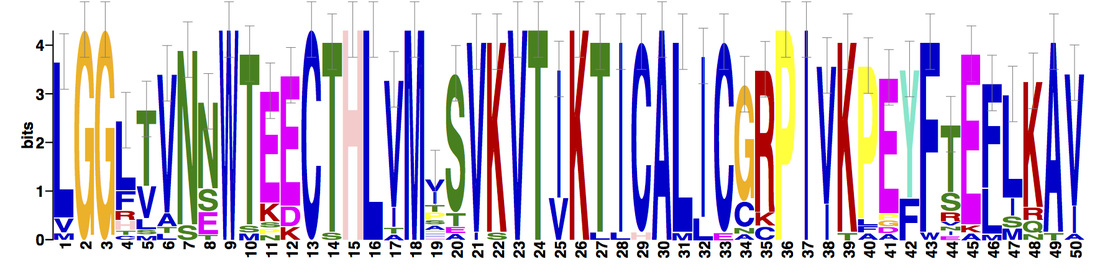
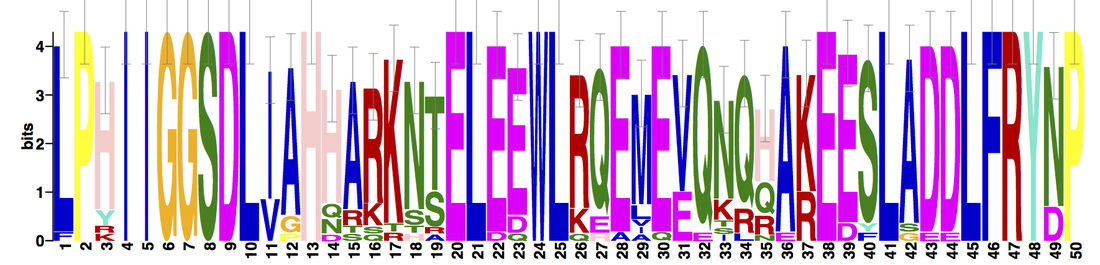
Three protein motifs were identified using MEME. Motifs were found to be conserved across all protein homologs.
|
Discussion
|
The NBN protein contains FHA, BRCT, and Nbs1_C domains. Results from Pfam, SMART, and Interpro report one of each domain while recent research has suggested the human NBN protein actually contains two, consecutive BRCT domains. All domains function in DNA damage repair, further supporting the primary function of the NBN protein. Additionally, the degree of domain and motif conservation across various organisms indicates their importance for organism viability.
NBN domains are highly conserved among vertebrates; particularly, mammalian and avian homologs, who display very little variation. Drosophila melanogaster (fruit fly), a potential inveterate model organism, lacks the Nbs1_C domain, but is still known to interact with Mre11 and the MRN complex [6]. Three motifs were found within the NBN protein homologs. Motifs 1 and 3 are positioned within the BRCT and Nbs1_C domains, respectively. Based on recent research and its downstream location from motif 1, motif 2 may further supports the potential existence of a second BRCT domain. |
|
References:
Figure 1. http://smart.embl-heidelberg.de/smart/show_motifs.pl?ID=NBN_HUMAN&DO_PFAM=DO_PFAM&INCLUDE_SIGNALP=INCLUDE_SIGNALP Figure 2. http://pfam.xfam.org/protein/nbn_human Figure 3. http://www.ebi.ac.uk/interpro/entry/IPR016592/proteins-matched Figures 4-6. http://www.ebi.ac.uk/interpro/entry/IPR016592/proteins-matched [1] Mullan, L. (2004). Domains and motifs – proteins in bite-sized chunks. BRIEFINGS IN BIOINFORMATICS, 5(1), 71-74. Retrieved February 28, 2015, from http://bib.oxfordjournals.org [2] Pawson, T. (2003). Assembly of Cell Regulatory Systems Through Protein Interaction Domains. Science, 445-452. Retrieved March 3, 2015, from http://web.b.ebscohost.com/ehost/detail/detail? [3] InterPro. (n.d.). Retrieved March 21, 2015, from http://www.ebi.ac.uk/interpro/entry/IPR000253?q=FHA domain [4] InterPro. (n.d.). Retrieved March 21, 2015, from http://www.ebi.ac.uk/interpro/entry/IPR001357?q=brct [5] InterPro. (n.d.). Retrieved March 21, 2015, from http://www.ebi.ac.uk/interpro/entry/IPR013908?q=nbs1_c [6] Gao, G., Bi, X., Chen, J., Srikanta, D., & Rong, Y. (2009). Mre11-Rad50-Nbs complex is required to cap telomeres during Drosophila embryogenesis. Proceedings of the National Academy of Sciences, 10728-10733. Retrieved March 24, 2015 from http://www.ncbi.nlm.nih.gov/pubmed/19520832?dopt=AbstractPlus&tool=FlyBase. |