This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison
|
Modifications
What are post-translational modifications?
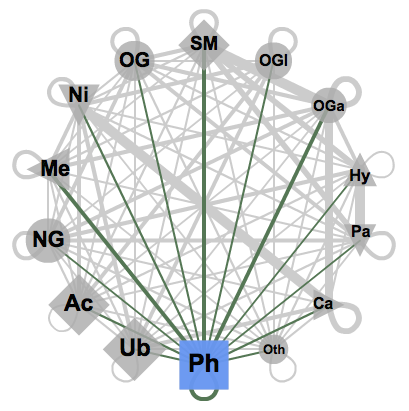
After a gene is transcribed and translated into a protein, it holds the potential to receive post-translational modifications (PTMs). As seen in Figure 1, there are a variety of PTMs including phosphorylations, gylcosylations, acetylations, ubiquitination, metylations, etc. Attached to specific amino acids side chains and peptide linkages of proteins, PTMs are chemical protein modifications associated with the regulation of activity, localization and interaction with other cell parts, such as proteins, nucleic acids, lipids and cofactors [1]. The presence of PTMs allow for the dramatic increase of protein interactions, functions, and ultimately the complexity and diversity of the proteome as a whole. Their continued study and understanding will open doors in research of cell biology while holding the potential to aid in the treatment and prevention of various disorders [1]. |
NBN Post-Translational Modifications
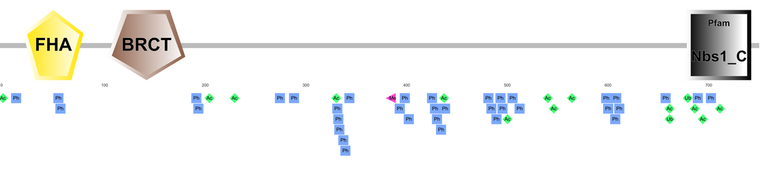
NetPhos 2.0 is a server that produces predictions of phosphorylation sites (serine, threonine and tyrosine) in eukaryotic proteins [3]. Differing from PTMcode, NetPhos predicted 56 human NBN phosphorylation sites; 39 serine, 14 threonine, and 3 tyrosine. Although many of the predictions possess a high phosphorylation potential, indicating a high likelihood that the amino acid is indeed phosphorylated, many other sites barely pass the threshold. This variation in confidence could explain the discrepancy between PTMcode and NetPhos regarding the number of phosphorylations on the human NBN protein.
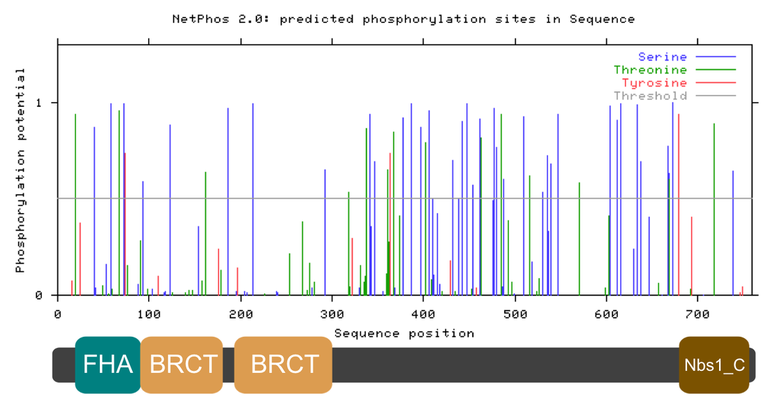
Figure 3. Phosphorylation sites of the human NBN protein according to predictions of NetPhos 2.0. The phosphorylation potential, or confidence score, is displayed for each predicted phosphorylation site. The key in the top right corner can be used to identify what amino acids where phosphorylated at specific sites along the NBN protein.
|
Discussion
|
After comparing results from both PTMcode and NetPhos, numerous PTMs were identified. Although acetylations, methylations and ubiquitinations were also seen, phosphorylations seemed the be the predominate PTM of the human NBN protein.
Taking a closer look, a majority of predicted phosphorylation sites lay between the BRCT and Nbs1_C domains, an area that currently possess no known domains. Of the predicted sites, serine seems to be most frequently phosphorylated, followed by threonine and tyrosine, respectively. |
|
References:
Figure 1. http://ptmcode.embl.de Figure 2. http://ptmcode.embl.de Figure 3. http://www.cbs.dtu.dk/services/NetPhos/ [1] Overview of Post-Translational Modifications (PTMs). (n.d.). Retrieved April 29, 2015, from https://www.lifetechnologies.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/overview-post-translational-modification.html [2] PTMcode 2. (2012). Retrieved May 2, 2015, from http://ptmcode.embl.de [3] NetPhos 2.0 Server. (Jan. 25 2015). Retrieved May 2, 2015, from http://www.cbs.dtu.dk/services/NetPhos/ |