This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison
|
Model Organisms
What are model organisms?
When an organism is used to study specific biological processes, they are considered a model organism. These animals, plants, and microbes possess simplified systems that are accessible and manipulatable. More often than not, they are chosen for their fast generation time, low complexity, and cost efficiency [1]. Because it is often unethical and impractical to carry out research on humans, utilizing model organisms to study human mutations has increased understanding, drug development and treatment options surrounding a variety of diseases. How do we find model organisms that are used to study specific diseases? Model organism databases. These systems store and provide genetic and genomic data of model organisms, such as yeast, worms, flies and mice [2]. When searching for organisms who possess a gene or protein of interest, results often provide mutant strains and their resulting phenotypes. If consistent with humans, these mutant phenotypes can infer potential model organisms. Results of mutant NBN strains from various model organism databases are provided below. Potential Model Organisms for NBS
|
Techniques to Obtain Mutant Strains
|
CRISPR-Cas 9
RNAi
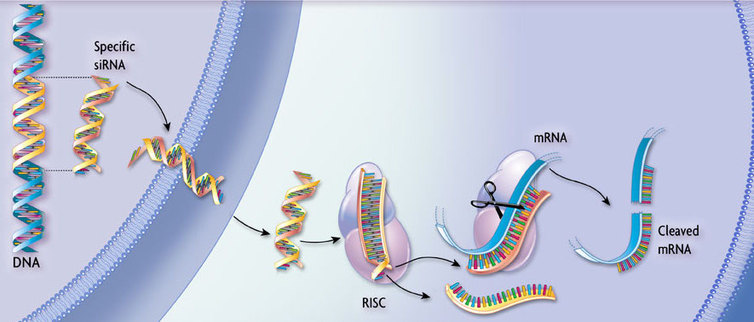
RNA interference (RNAi) is a technique to knockdown, or silence, specific gene activity. This process is completed by destroying mRNAs, therefore preventing gene translation and overall protein production. mRNA is destroyed due to the introduction of double-stranded RNA molecules into the cell. These molecules are then fragmented by dicer, an enzyme responsible for cleaving double-stranded RNA. Once cleaved, the RNA fragments find complementary mRNA sequences, bind, and attract the RNAi machinery (dicer), leading to mRNAs' fragmentation as well [4](Figure 1).
|
Discussion
|
Identifying appropriate model organisms is an important step in understanding Nijmegen breakage syndrome. Drosophila melanogaster and Mus musculus were the only organisms to have reported mutant phenotypes due to mutations in NBN. Because a majority of mutant phenotypes experienced by both organisms were consistent with human NBS patients, both would pose as efficient representations of the disease. However, because fruit flies are more cost effective, utilizing their small size, fast generation time, and low costs would make them a better option. On the contrary, fruit flies would not be the best choice to model cancer associated with NBS. For research aimed at cancer development, mice, who possess a greater probability of developing malignancies during their lifetime, would be a better representation.
Additionally, utilizing the naked mole-rat as a model organism may lead to ground breaking discoveries in NBS cancer research. These rodents have an astonishing low susceptibility to cancer. Because cancer is the leading cause of death in NBS patients, studying NBN mutations in this organism may be extremely beneficial. There is rarely only one model organism used to study a human disease. As we see with NBS, each organism has its own benefits and drawbacks. When carrying out research we first look to the organism that is a best representation of what we are studying. Drosophila melanogaster is a logical choice for studying NBN at a molecular level, however, if we choose to study more longterm outcomes of NBS, such as cancer, Mus musculus becomes the better option. |
|
References:
Video 1. https://www.youtube.com/watch?v=2pp17E4E-O8 Figure 1. http://www.alnylam.com/Files/Media_Kit/ALNY-ILLUS_01_RNAiNaturalProcess.jpg [1] Using Model Organisms to Study Health and Disease. (2012). Retrieved March 27, 2015 from, http://www.nigms.nih.gov/Education/Pages/modelorg_factsheet.aspx [2] GMOD Main Page. (2011). Retrieved March 27, 2015, from http://gmod.org/wiki/Main_Page [3] SITNFlash. (2014, July 13). CRISPR: A game-changing genetic engineering technique. Retrieved March 27, 2015, from http://sitn.hms.harvard.edu/flash/2014/crispr-a-game-changing-genetic-engineering-technique/ [4] RNA Interference Fact Sheet. (2012). Retrieved March 27, 2015 from, http://www.nigms.nih.gov/News/Extras/RNAi/Pages/factsheet.aspx |